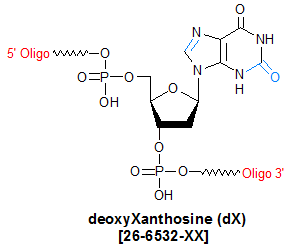
Modification : deoxyXanthosine
Catalog Reference Number
Category
Modification Code
5 Prime
3 Prime
Internal
Molecular Weight (mw)
Extinction Coeficient (ec)
Technical Info (pdf)
Absorbance MAX
Emission MAX
Absorbance EC
26-6532
Structural Studies
[dX]
Y
Y
Y
330.19
15.5
PS26-6532.pdf
-
-
-
| Catalog No | Scale | Price |
| 26-6532-05 | 50 nmol | $274.00 |
| 26-6532-02 | 200 nmol | $274.00 |
| 26-6532-01 | 1 umol | $357.00 |
| 26-6532-03 | 2 umol | $535.00 |
| 26-6532-06 | 5 umol | $1,606.50 |
| 26-6532-10 | 10 umol | $2,853.00 |
| 26-6532-15 | 15 umol | $3,566.00 |
| Discounts are available for deoxyXanthosine! |
| Modification* Discount Price Structure |
|
1 site/order
|
List price
|
|
2 sites/order
|
10% discount
|
|
3 sites/order
|
20% discount
|
|
4 sites/order
|
30% discount
|
|
5-9 sites/order
|
50% discount
|
|
10+ sites/order
|
60% discount
|
|
*Exceptions apply
|
Deoxyxanthosine (dX) is classified as an oxidized nucleoside, and is believed to be produced in cellular DNA via oxidative deamination of dG. Because dX has some potential to pair with all four natural bases, it has been explored for its utility as a universal base (1). In the mid-1990s, the base-pairing affinity of dX for 2,4-diaminopyrimidine (py(DAD)), as well as the ability of several DNA polymerases to incorporate py(DAD) opposite dX, was studied in order to explore the potential of using the dX:py(DAD) non-standard base pair to expand the genetic alphabet (2). The authors of that study found that DNA polymerase incorporated py(DAD) opposite dX with high efficiency that was pH-independent.
Because dX is an oxidized nucleoside, there is also considerable interest in using it in studies of oxidative DNA damage and associated repair mechanisms. In particular, because nitric oxide (NO) is known to react with guanine to produce xanthine (X), dX-modified oligonucleotides have been used to study the biochemical role of X in NO-mediated mutagenesis (3). dX can potentially pair with C (leading to no mutation) and mispair with T (leading to G-to-A transitions). As a single-base lesion, dX is preferentially removed by the base excision repair (BER) mechanism when it is paired opposite C, and the native guanine base restored.
References
1. Eritja, R., Horowitz, D.M., Walker, P.A., Ziehler-Martin, J.P., Boosalis, M.S. Synthesis and properties of oligonucleotides containing 2’-deoxynebularine and 2’-deoxyxanthosine.
Nucleic Acids Res. (1986),
14: 8135-8153.
2. Lutz, M.J., Held, H.A., Hottinger, M., Hubscher, U., Benner, S.A. Differential discrimination of DNA polymerase for variants of the non-standard nucleobase pair between xanthosine and 2,4-diaminopyrmidine, two components of an expanded genetic alphabet.
Nucleic Acids Res. (1996),
24: 1308-1313.
3. Wuenschell, G.E., O’Connor, T.R., Termini, J. Stability, miscoding potential, and repair of 2’-deoxyxanthosine in DNA: implications for nitric oxide-induced mutagenesis.
Biochemistry (2003),
42: 3608-3616.
- deoxyXanthosine
