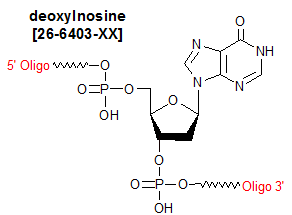
Modification : Inosine deoxy
Catalog Reference Number
Category
Modification Code
5 Prime
3 Prime
Internal
Molecular Weight (mw)
Extinction Coeficient (ec)
Technical Info (pdf)
Absorbance MAX
Emission MAX
Absorbance EC
26-6403
Duplex Stability
[dI]
Y
Y
Y
314.19
7.5
PS26-6403.pdf
-
-
-
| Catalog No | Scale | Price |
| 26-6403-05 | 50 nmol | $17.00 |
| 26-6403-02 | 200 nmol | $17.00 |
| 26-6403-01 | 1 umol | $24.00 |
| 26-6403-03 | 2 umol | $38.00 |
| 26-6403-06 | 5 umol | $76.00 |
| 26-6403-10 | 10 umol | $104.00 |
| 26-6403-15 | 15 umol | $130.00 |
Deoxyinosine (dI) is classified as a nucleotide base analog; it is structurally similar to guanosine, but is missing the 2-amino group. Because it is able to form two hydrogen bonds with each of the four natural nucleotide bases (1), it is often used by researchers as a "universal" base meaning that it can base pair with all the naturally-occurring bases--in synthetic oligos. dI typically is substituted for the nucleoside at the third ('wobble') position of codons, in order to reduce the complexity of mixed oligo PCR primers/hybridization probes needed to deal with degenerate codons in the target DNA (2, 3). However, it is important to remember that dI does not base pair equally well with the naturally-occurring bases, with the order of thermodynamic stability being I-C >: I-A > I-G ~ I-T. Thermodynamic stability of inosine-containing duplexes is also affected by neighboring bases (4). Consequently, when using dI as an alternative to mixed-base degeneracy at a particular oligo position, keep in mind that the above base-pairing bias may lead to differences in the oligo's priming or hybridization efficiency in the corresponding degenerate regions of the target. Because the effect could be more pronounced when dI is at the 3'-position, it may be advisable to use primers with and without dI at the 3'-end, in order to maximize diversity of PCR products (5).
References
1. Oda, Y, Uesugi, S., Ikehara, M., Kawase, Y., Ohtsuka, E. NMR studies for identification of dI:dG mismatch base-pairing structure in DNA.
Nucleic Acids Res. (1991),
19: 5263-5267.
2. Liu, H., Nichols, R. PCR amplification using deoxyinosine to replace entire codon and at ambiguous positions.
Biotechniques. (1994),
16: 24-26.
3. Ohtsuka, E., Matsuki, S., Ikehara, M., Takahashi, Y., Matsubara, K. An alternative approach to deoxynucleotides as hybridization probes by insertion of deoxyinosine at ambiguous codon positions.
J. Biol. Chem. (1985),
260: 2605-2608.
4. Martin, F.H., Castro, M.M., Aboul-ela, F., Tinoco, I. Base pairing involving deoxyinosine: implications for probe design.
Nucleic Acids Res. (1985),
13: 8927-8938.
5. Ben-Dov, E., Shapiro, O.H., Siboni, N., Kushmaro, A. Advantage of Using Inosine at the 3' Termini of 16S rRNA Gene Universal Primers for the Study of Microbial Diversity.
Appl. Environ. Microb. (2006), 72: 6902-6906.
- Inosine deoxy (dI)
